Circular oligonucleotides are an emerging class of nucleic acid therapeutics (NATs) because of their resistance to exonucleases and their reduced immunogenicity.[1] Wesselhoeft et al., demonstrated that exogenous circRNA did not activate Toll-like Receptors (TLRs) nor Retinoic acid-Inducible Gene I (RIG-I), provided that the circRNA was meticulously purified to remove all linear RNA fragments. In addition, the duration of protein translation was extended significantly compared to its linear counterpart.[2] Also shown was the compatibility of circRNAs to be packaged and delivered by Lipid Nanoparticles (LNPs) into tissue with concomitant expression of the translated protein.
Other labs have focused on the use of circular ASOs to knockdown gene expression such as the labs of Dongsheng Liu and Yuanchen Dong. Using DNA ligase, they made large, circular single-stranded DNA ASOs, 71 and 132 nucleotides in length, to target single, and multiple mRNA transcripts, respectively. Their recently published paper demonstrated that circular, single-stranded DNA could activate RNase H and successfully knockdown target gene expression.[3] However, long circASOs were necessary for the circASO to hybridize to the target mRNA without undue torsional stress. For the 71nt circASO, only 22 bases actually hybridized to the target sequence, the rest being essentially a large spacer. This is, of course, problematic due to the expense and difficulty of the synthesis and purification, especially upon scaling up.
There have been some creative ways to get around this problem. Xinjing Tang’s lab circularized a 26mer ASO using a peptide linker that was a substrate to cathepsin B.[4] By doing so, the circASO would remain inactive until it was cleaved within the endosome. They demonstrated that the short 26 nt circASO successfully knocked down the targeted mRNA but only in cells that expressed cathepsin B. However, the synthesis was still quite involved given the peptide conjugation, activation and final circularization of the ASO.
Arguably, the most practical and creative solution came out of Sudhir Agrawal’s lab.
Agrawal had explored ligating 3’-to-3’ and 5’-to-5’ linked linear oligonucleotides, which generated oligos with two 5’ termini and two 3’ termini respectively. Unexpectedly, these had very different immunostimulatory properties with the 3’-3’ linked ASOs highly immunogenic whereas the 5’-5’ linked ASO were minimally so. In addition, 5’-5’ linked ASO were approximately twice as potent in knocking down its target mRNA despite the 5’-5’ and 3’-3’ sequences being the same.[5] However, the problem with this design was two-fold: first, the length of the 5’-5’ linked ASO is twice as long as the standard ASO, raising synthesis and purification costs. Second, most exonuclease activity is from the 3’ terminus, limiting its stability in plasma. Agrawal’s creative solution was a new design where only the last ~6-10 bases were 5’ orientation, however, these bases were complementary to the last ~6-10 bases of the other 3’ terminus.[6] The structure of this “transient cyclic structured oligonucleotide” (CSO) is shown in Fig. 1.
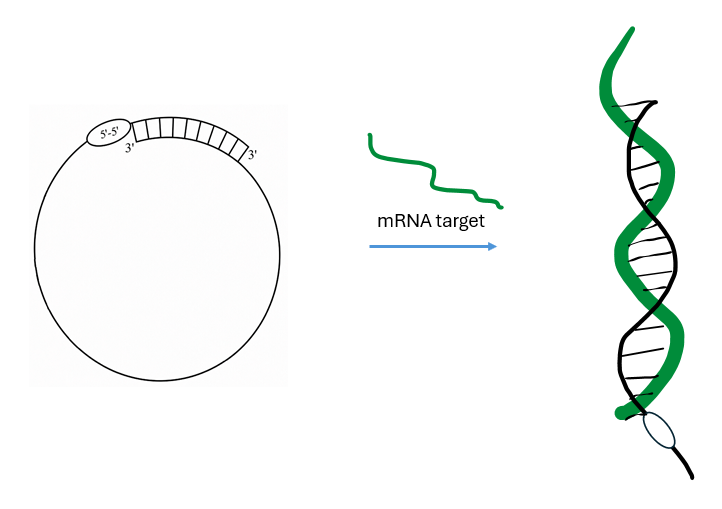
Fig. 1 Shown on the left is the schematic of the transient cyclic structured ASO. By having the reverse bases after the 5’-5’ linkage complementary to the 3’ bases on the other end of the oligo, both the 3’ ends are now protected from 3’ exonucleases by virtue of being double-stranded.
By balancing the melting temperature (Tm) of the probe-target duplex to be approximately 20 °C higher than that of the 3’ circularizing domain, in the presence of the target mRNA, the circularized oligo will open up and hybridize to the target, recruiting RNase H. The specific melting temperatures suggested by Agrawal are 42-55 °C for the circularizing domain and 60-75 °C for the probe-target duplex.[6]
The advantages of the CSO design are:
- Reduced immunogenicity (innate immune responses are very sensitive to the single-stranded 5’ terminus and this structure has none)
- Enhanced stability (exonucleases won’t touch it)
- Improved target specificity and mismatch discrimination
The improved target specificity comes from the equilibrium between the circularized and linear forms of the ASO as shown in Fig. 2.
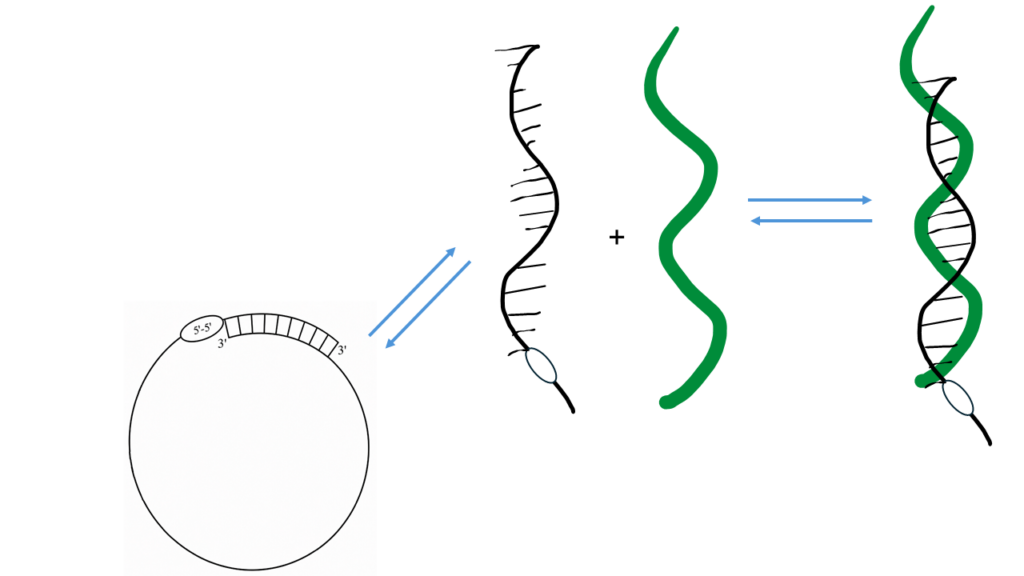
Fig. 2 The stability of the circularized structure drives the equilibrium away from the duplexed structure. This leads to higher sensitivity to base mismatches in the ASO. By reducing or lengthening the circularizing domain, that balance can be finely tuned to optimize specificity.
This is analogous to the behavior of molecular beacons where there is an equilibrium between the stem-loop structure and that of the hybridized duplex which leads to higher mismatch discrimination. This increased mismatch discrimination is borne out by experiment as well as theory – which is generalizable to all conformationally constrained probes.[7]
There are several advantages to this design, the first of which is its simplicity. The CSO ASO is synthesized using standard phosphoramidites and SPOS chemistry but using 5’ phosphoramidites (otherwise known as reverse phosphoramidites) for the last 6-10 bases. This makes the CSO:
- Simple to manufacture without the need for splints or ligases
- You can make a short, circularized oligo that still hybridizes to the target because the circularization is reversible in the presence of the target
- No exposed 3’ or 5’ termini – which greatly reduces immunogenicity and while increasing nuclease resistance
- Improved target specificity – while having a high Tm duplex with the target.
- Improved potency over a linear ASO with the same targeting sequence
Agrawal then added an interesting twist to the design – one of the strands of the circularization domain was RNA rather than DNA, which he named a Cyclic Prodrug Oligonucleotide (CPO). Upon release into the cytoplasm, RNase H cleaves it, linearizing it to its active form. Data in reference [6] showed this design approximately an order of magnitude more potent than the standard CSO design (which in turn was an order of magnitude more potent than the linear ASO). Yet another design of therapeutic potential is the ring-shaped oligonucleotide (RSO) shown in Fig. 3.
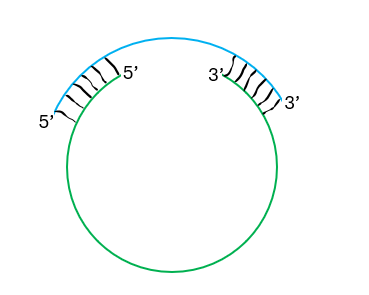
Fig. 3 Shown here is a schematic of a ring-shaped oligonucleotide (RSO) with the functional domain in green and the circularizing domain in blue. The circularizing domain can be either DNA or RNA depending upon whether the ring is opened by hybridization to the target or assisted by RNase H activity.
In this case a large domain (shown in green) is the active therapeutic oligo, such as a single-guide RNA, held in circularized secondary structure by an oligo.
In conclusion, there are a great variety of different approaches to explore in this new class of cyclic therapeutic oligos, covalently circularized through a phosphodiester or phosphorothioate linkage, covalently circularized using a cleavable linker, such as a peptide or finally transiently circularized (which may or may not utilize RNase H to facilitate linearization). There are many options to explore and it will be quite interesting to see which design, or designs, make it to the clinic.
References
[1] Zhen X, Mahmoudi M, Lan X, Huang G, Tao W. Unlocking the potential of engineered circular RNA therapeutics. Med X. 4(1):7 (2026). doi: 10.1007/s44258-026-00079-5.
[2] RA Wesselhoeft, PS Kowalski, FC Parker-Hale, Y Huang, N Bisaria, DG Anderson, RNA Circularization Diminishes Immunogenicity and Can Extend Translation Duration In Vivo. Molecular Cell 74, 508–520 (2019).
[3] Yufan Pan, Xin Li, Chenyou Zhu, Mingyan Wang, Yuanyuan Wu, Rui Xu, Jun Wu, Ruofan Chen, Yifan Jiang, Baolei Tian, Yuanchen Dong, and Dongsheng Liu, Programmable Single-Stranded Circular Antisense Oligonucleotides for Multitarget Gene Therapy. JACS Au, vol 5, 3555−3564 (2025).
[4] Cathepsin B-activatable cyclic antisense oligonucleotides for cell-specific target gene knockdown in vitro and in vivo. Zhongyu Wang, Xinli Fan, Guanqun Mu, Xiaoran Zhao, Qian Wang, Jing Wang, and Xinjing Tang, Molecular Therapy: Nucleic Acids, Vol 33, 548-558 (2023).
[5] Bhagat, L., Putta, M. R., Wang, D., Yu, D., Lan, T., Jiang, W., Sun, Z., Wang, H., Tang, J. X., la Monica, N., Kandimalla, E. R., & Agrawal, S. Novel oligonucleotides containing two 3’-ends complementary to target mRNA show optimal gene-silencing activity. Journal of Medicinal Chemistry, 54(8), 3027–3036 (2011)
[6] Agrawal, S. Transient cyclic structured oligonucleotide designs for therapeutic applications. Current Protocols, 6, e70319, (2026). doi: 10.1002/cpz1.70319
[7] Bonnet G, Tyagi S, Libchaber A, Kramer FR. Thermodynamic basis of the enhanced specificity of structured DNA probes. Proc Natl Acad Sci, 96(11):6171-6176 (1999). doi: 10.1073/pnas.96.11.6171.