Background
Ligases are ubiquitous. They are found in every living organism and are required for cell division – stitching together the Okazaki fragments produced during DNA replication and maintaining genetic integrity by performing the final step in every DNA repair pathway – the crucial ligation step – which is the formation of a phosphodiester linkage between a 5’-phosphorylated nucleic acid strand and the 3’ hydroxyl of another.
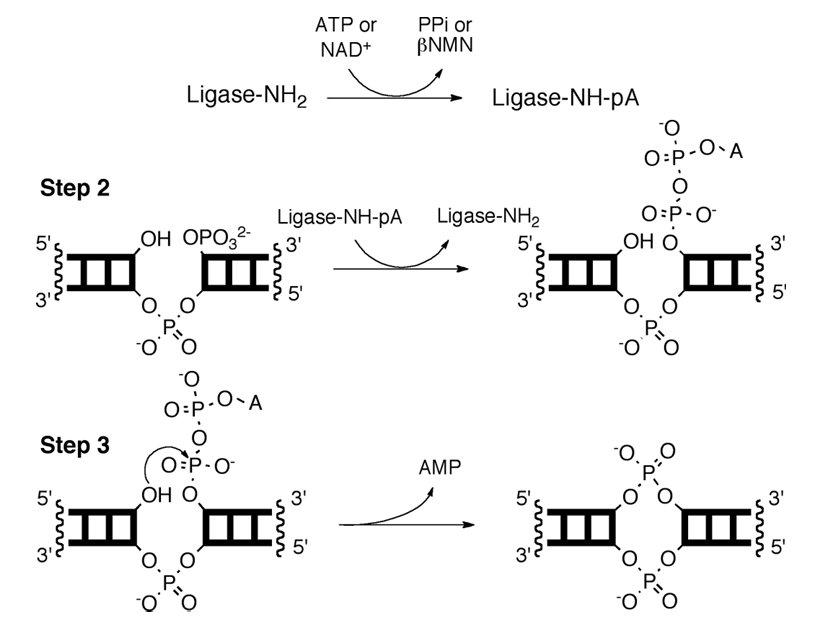
Ligases are divided into two broad groups – DNA and RNA ligases. RNA ligases are further divided into single- and double-stranded ligases where single-stranded ligases are template-free and double-stranded ligases are template-dependent. Given the nature of DNA in the cells, DNA ligase substrates are always double-stranded except in some viruses. Shown in Fig. 1 is the mechanism of ligation. First, the ligase is charged by either NAD+ or ATP to adenylate a lysine in the active site of the enzyme. The adenylated enzyme then transfers the adenylyl group onto the 5’ phosphate. Finally, the 3’-hydroxyl reacts with the 5′-adenylyl pyrophosphate, forming the phosphodiester linkage with AMP as the leaving group. Note, only ligases derived from bacteria use NAD+ as the co-substrate in the reaction.
Template-dependent convergent synthesis
One of the most useful aspects of ligases is that, unlike polymerases, their tolerance to both sugar and base modifications is quite high. This allows standard phosphoramidite chemistry to be used to make highly modified, short oligos and then use either a template[1] or another oligo to act as the initial template,[2] to ligate the short oligos together. Shown in Fig. 2 (A) is a schematic of template-dependent convergent synthesis.
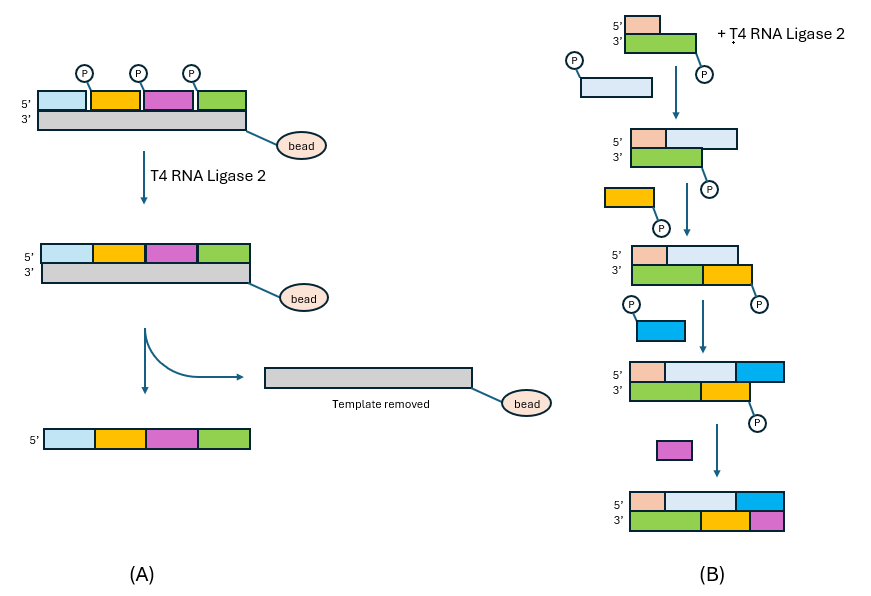
In this case, the oligos are annealed and then co-ligated on the template in a single step. The template is biotinylated and attached to a streptavidin-coated magnetic bead. After the ligation is completed, the beads are pulled down with a magnetic separation rack and repeatedly washed to allow the fast and simple removal of the excess unligated oligos. The final ligated, full-length product is released from the template-bead conjugate by briefly raising the pH to 12. Shown in Fig. 2 (B) is the schematic of a template-dependent convergent synthesis where consecutive ligations are performed to ‘boot-strap’ to a final, double-stranded siRNA.[2] In this case, the ‘template’ is part of the final siRNA product unlike in Fig. 2 (A) where the template is removed after the ligation is completed. The advantage of a convergent synthesis such as this is measured in Process Mass Intensity (PMI). PMI is the ratio of the mass of raw material that goes into the process divided by the mass of the Active Pharmaceutical Ingredient (API) that is produced.
A recent paper evaluated eight oligonucleotide APIs currently in development. Their calculated PMIs range from 3035−7023.[3] High PMI values translate to high costs and an extremely expensive drug. Convergent syntheses, however, can reduce the PMI substantially. First, the short, starting oligos used in convergent syntheses will often not require any purification simply because they are short and fewer impurities will build up within the sequence. Second, the addition of 5’ phosphate is the final step of the solid phase oligonucleotide synthesis and because of this, capped failure sequences cannot ligate. Finally, the problematic n-1mers will either not be adjacent to the 3’-hydroxyl or there will be a mismatch that can allow for discrimination between the full-length oligo and the n-1mers. This is borne out by results shown in Fig. 3 where the same siRNA was synthesized three different times using either four, five or six short oligos (corresponding to two, three or four ligation events). Note the total percentage of impurities in the starting oligos goes down as the number of oligos used to make the siRNA increases. More importantly to note is that with an increasing number of ligation events, a higher percentage of impurities were rejected from the FLP – with impurities of less than 2% in the final siRNA synthesized with four ligation events without purification other than ultrafiltration.[2]
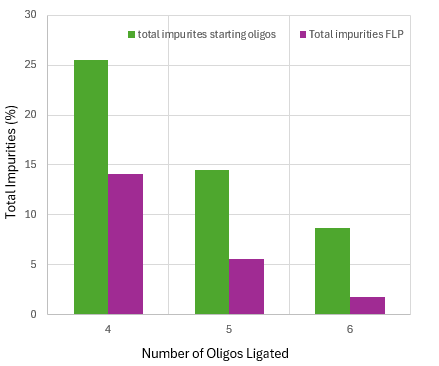
Fig. 3 Plot of percent total impurities of the starting oligos used without purification versus percent total impurities found in the full-length, siRNA product. The same siRNA was synthesized three times using either four, five or six small starting oligos (corresponding to two, three or four ligation events). Based upon supplemental data from Paul, S., et al., Convergent Biocatalytic Mediated Synthesis of siRNA, ACS Chem. Biol. 18, 2183−2187 (2023).
The success of this strategy depends upon a careful choice of how the siRNA sequence is broken up into short oligos as well as conditions to optimize ligation efficiency. Choosing the best ligase for the application, of course, is crucial to success. To allow this discrimination of n-1mers from the full-length oligo and to prevent annealing of mismatched oligos, thermostable versions of the T4 RNA Ligase 2, among others, are typically used.
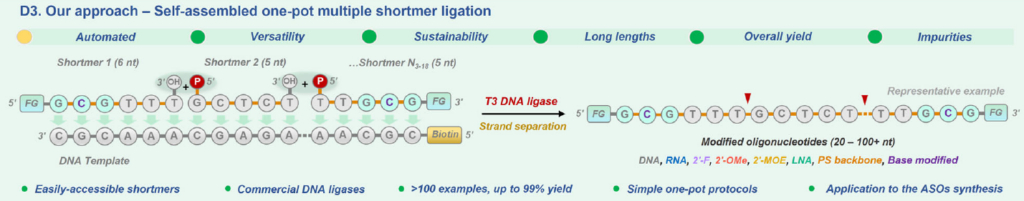
The Hollenstein lab did a very thorough investigation of DNA template-based synthesis (Fig. 4) evaluating not just a variety of DNA ligases – e.g. T3, T4, T7 – they also looked at the effect of molecular crowding using PEG, weak denaturants like DMSO, and alternative metal salts, such as Mn2+ among others, to optimize ligation efficiency of highly modified ASOs. The modifications included 2’-F, 2’-OH, 2’-MOE, 2’-OMe, and LNA.[1] To confirm the robustness of the protocol, they successfully synthesized Fomivirsen, a fully-phosphorothioate backbone ASO. Note, to insert a phosphorothioate linkage at the ligation site, a 5’-thiophosphate oligo is used, however only the Rp phosphorothioate enantiomer is produced which has substantially reduced nuclease resistance compared to the Sp enantiomer.

Use of Ligases for Production of CircRNA and Long sgRNAs
A recent paper that came out of Genentech/Roche described the production of sgRNA using T4 RNA Ligase 2.[4] Single-guide RNAs are long, modified RNAs for which the linear chemical synthesis by standard phosphoramidite is challenging, given the length of the RNA and the purity requirements. The 100mer was essentially broken in half into a 49 base and 51 base sequences. A 28 base DNA splint was used to bring the phosphorylated 51mer adjacent to the 3’-hydroxyl 49mer. After allowing the splint to hybridize to the other strands, T4 RNA Ligase 2 was added to ligate the strands together. (It should be noted that the T4 RNA Ligase 2 is rather promiscuous and can ligate DNA:RNA heteroduplexes). They were able to successfully use this convergent synthesis strategy on a tens-of-grams scale, with 3-4 fold higher yield and 10% higher purity.
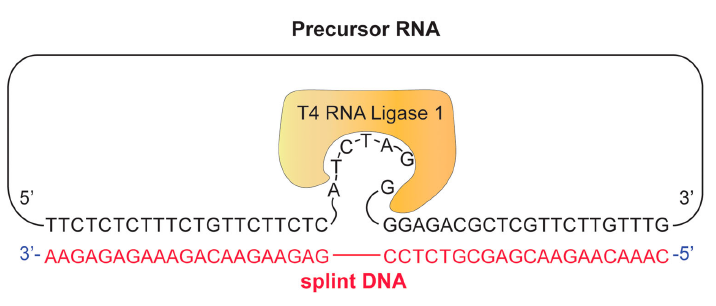
Another really interesting paper that has flown under the radar came out of a collaboration between the lab of Hak Kyun Kim at Chung Ang University and NuclixBio in Seoul.[5] Their goal was to efficiently produce circRNA. CircRNA is significantly more stable than linear RNA and is involved in a variety of cellular functions. If an circRNA contains an IRES in front of the coding sequence (CDS), the circRNA can be translated in vivo into proteins by the ribosomes for therapeutic uses.[5] Kim and NuclixBio were able to successfully circularize 1.3 kb CDSs up to a 4.5 kb CDS with a whopping 70% efficiency for the 1.3 kb transcript and a very respectable 15% efficiency for the 4.5 kb transcript. Interestingly, they did not use T4 RNA ligase 2, but rather T4 RNA ligase 1, which normally just works on ligating single-stranded RNAs. They did so by designing the splint to bulge out 7 bases as shown in Fig. 5. The authors found that even a 26mer could efficiently be used to facilitate circularization, but only if diaminopurine (DAP) was used to increase the stability of the duplex. Without the DAP, a 42mer was required to achieve the same circularization efficiency. It will be interesting to see if a similar strategy can be used to ligate long, single-stranded RNAs such as pegRNA.
Here at SynOligo Biotechnologies, we work with our customers from discovery to the pre-clinical scale, including cell-based screening assays for potency and selectivity of siRNAs and ASOs, including assessment of toxicity and immune response activation in cells. We endeavor to minimize our customer’s costs while maintaining the highest standards of quality. We feel that the use convergent synthesis strategies will be utilized more and more frequently in the field as the demand for therapeutic, highly pure and affordable RNA longmers continues to grow.
References
[1] Sabat, N., Stämpfli, A., Hanlon, S. et al., Template-dependent DNA ligation for the synthesis of modified oligonucleotides. Nat Commun 15, 8009 (2024). https://doi.org/10.1038/s41467-024-52141-8
[2] Paul, S., et al., Convergent Biocatalytic Mediated Synthesis of siRNA, ACS Chem. Biol. 18, 2183−2187 (2023).
[3] Andrews, B.I. et al., Sustainability Challenges and Opportunities in Oligonucleotide Manufacturing, J. Org. Chem. 86, 49−61(2021).
[4] Bigatti, M., et al., Development of a Broadly Applicable Enzymatic Ligation Process for the Production of Single Guide RNAs, Org. Process Res. Dev. 29, 5, 1228–1236 (2025).
[5] Yoon-Seob Kim, Do-Hyung Kim, Daegi An, Younghyun Lim, Young-Jin Seo, Hak Kyun Kim & Ho-Young Kang, The RNA ligation method using modified splint DNAs significantly improves the efficiency of circular RNA synthesis, Animal Cells and Systems, 27:1, 208-218 (2023)
